RNAi Database sample wiki
Contents
Sample of the RNAi Database Dashboard
Overview
- RNA interference (RNAi), is a technique in which exogenous, double-stranded RNAs (dsRNAs) that are complimentary to known mRNAs, are introduced into a cell to specifically destroy that particular mRNA, thereby diminishing or abolishing gene expression.
- This technology considerably bolsters functional genomics to aid in the identification of novel genes involved in disease processes and thus can be used for medicament and for delivery as therapeutics. Source
- RNA interference was known by other names, including post transcriptional gene silencing and quelling. Source
Effector RNA molecules
RNAi pathways are guided by small RNAs that include
SiRNA:
- Small interfering RNA (siRNA), sometimes known as short interfering RNA or silencing RNA, is a class of 20-25 nucleotide-long double-stranded RNA molecules.
- SiRNAs can also be exogenously (artificially) introduced into cells by various transfection methods to bring about the specific knockdown of a gene of interest. Source
miRNA:
- microRNAs (miRNA) are single-stranded RNA molecules of about 21–23 nucleotides in length, which regulate gene expression.
- miRNAs are encoded by genes from whose DNA they are transcribed but miRNAs are not translated into protein (non-coding RNA); instead each primary transcript (a pri-miRNA) is processed into a short stem-loop structure called a pre-miRNA and finally into a functional miRNA.
- Mature miRNA molecules are partially complementary to one or more messenger RNA (mRNA) molecules, and their main function is to down-regulate gene expression. Source
shRNA:
- A small hairpin RNA or short hairpin RNA (shRNA) is a sequence of RNA that makes a tight hairpin turn that can be used to silence gene expression via RNA interference.
shRNA uses a vector introduced into cells and utilizes the U6 promoter to ensure that the shRNA is always expressed. This vector is usually passed on to daughter cells, allowing the gene silencing to be inherited. The shRNA hairpin structure is cleaved by the cellular machinery into siRNA, which is then bound to the RNA-induced silencing complex (RISC). This complex binds to and cleaves mRNAs which match the siRNA that is bound to it. Source
Others:
- In addition to miRNAs and siRNAs, other innate RNAi effectors have been identified.
- One class of these is the Piwi-interacting RNAs (piRNAs). piRNAs seem to be uniquely expressed in the mammalian germline, particularly in the testes. The functional role of piRNAs is currently unclear, but a role in spermatogenesis is likely.
- A number of other small RNAs associated with RNAi have been identified in different species, including trans-activating siRNAs (tasiRNAs), studied in plants and nematodes, and small scan RNAs (ScnRNAs), found in Tetrahymena. Source
Cellular Mechanism
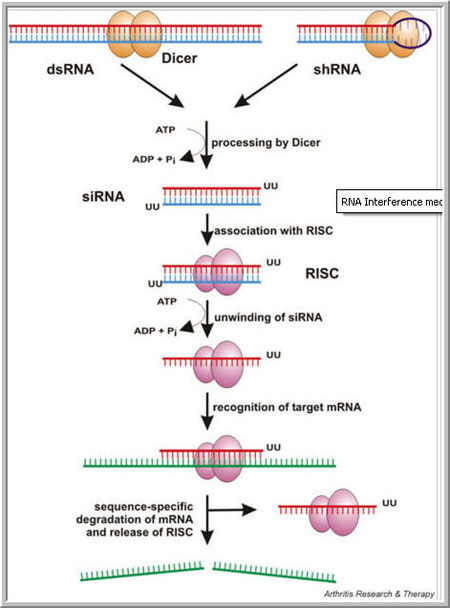
The RNA interference pathway: Long double-stranded RNA (dsRNA) or small hairpin RNA (shRNA) is processed by Dicer to form a small interfering RNA (siRNA), which associates with RNA-induced silencing protein complex (RISC) and mediates target sequence specificity for subsequent mRNA cleavage leading to gene silencing.
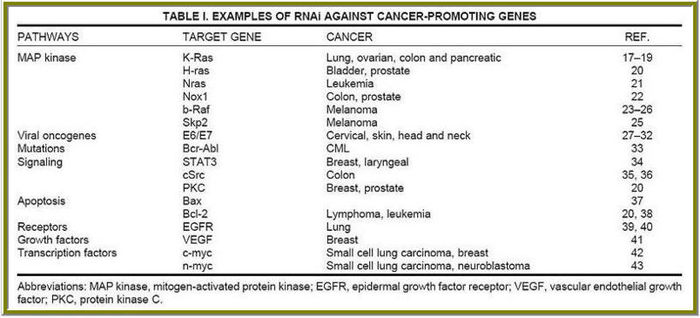
Cancer specific Targets
Intellectual property
Search Strategy
Effective search strategy ensures that all key words covering the technology are adequately covered. IPC class limitations help in removing the unwanted patents.
The search strategy was employed to extract patents disclosing RNAi agents against disease causing target genes. Table below summarizes an exemplary search strategy used to extract patents for BCL2, a target gene for curing cancer.
| S.No. | Concept | Search scope | Query | No. of Hits |
| 1 | RNAi | Claims,Title, Abstract | (siRNA*1 OR (short ADJ1 interfering ADJ1 RNA*1) OR (short ADJ1 interfering ADJ1 nucleic ADJ1 acid*1) OR (short ADJ1 interfering ADJ1 NA*1) OR siNA OR siNAs OR (si ADJ1 RNA*1) OR (Small adj1 interfering adj1 RNA*)OR miRNA or (microRNA* or micro adj1 RNA*)OR shRNA or (small adj1 hairpin adj1 RNA*)OR piRNA or (Piwi adj1 interacting adj1 RNA*)OR tasiRNAs or (trans adj1 activating adj1 siRNA*) OR ScnRNAs or (small adj1 scan adj1 RNA*)OR RNAi or(RNA adj1 interference)) | 12729 |
| 2 | IPC | a61k or c12q or c12n or c07k or c07h | 1688146 | |
| 3 | BCL2 | Full Patent Spec | (Bcl2 OR (Bcell ADJ1 CLL) OR (Bcell ADJ1 Lymphoma2 ) OR DKFZp781P2092) and Current IPC-R a61k or c12q or c12n or c07k or c07h | 3890 |
| 4 | BCL2 + RNAi +IPC | 1 AND 3 | 633 (293 unique records) |
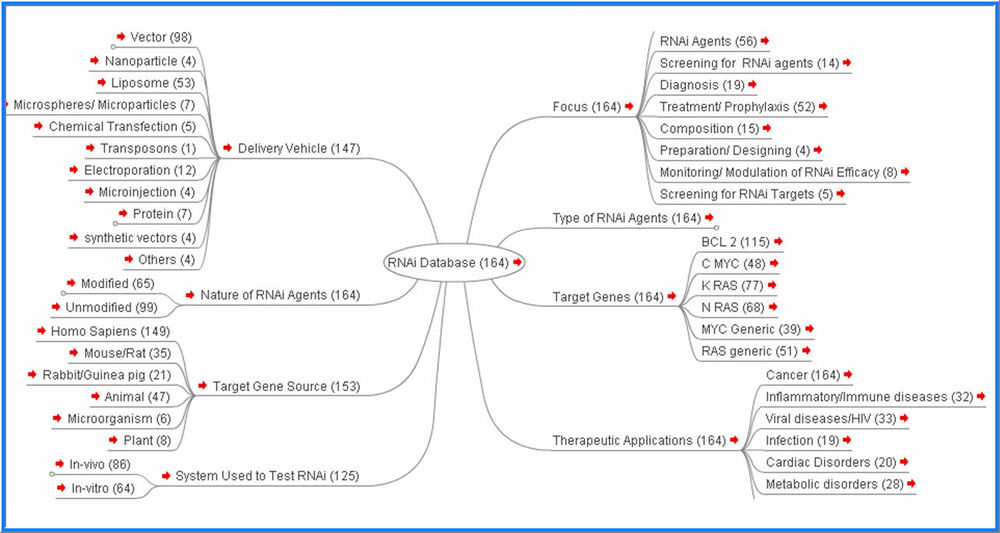
Interactive Taxonomy
Grouping specific elements into more general categories is conceptually easier and cleaner than is entertaining hundreds of specific elements separately. We categorize the information from analyzed patents and scientific literate into four levels of taxonomy which allow both “bird’s-eye view” and “in-depth views” of scientific technologies.
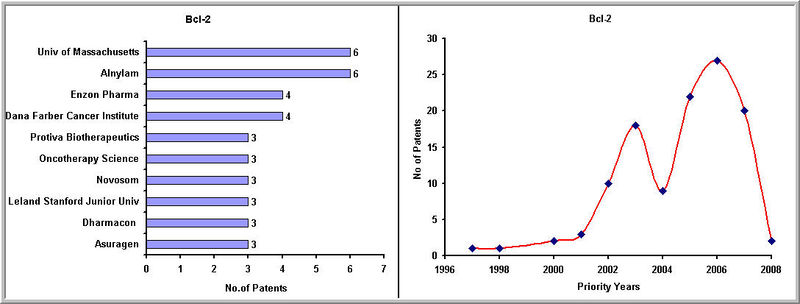
Top Players and Patenting Activity
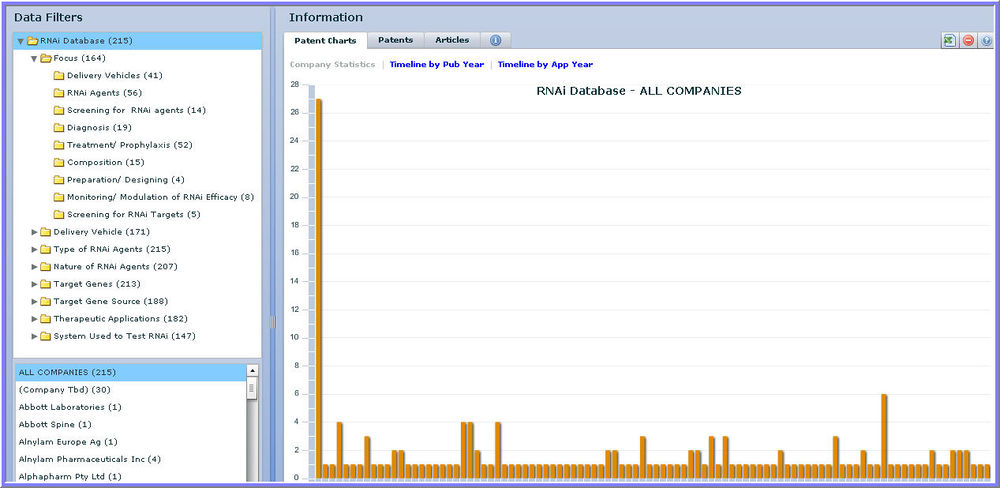
RNAi Database Dashboard
RNAi database dashboard:
- Categorize and organize large sets of product, patent, and scientific literature
- Analyze client’s competitive position and that of key competitors
- Identify licensing opportunities
- Make build-vs.-buy decisions in new product areas
- Predict product features based on technology and market trends
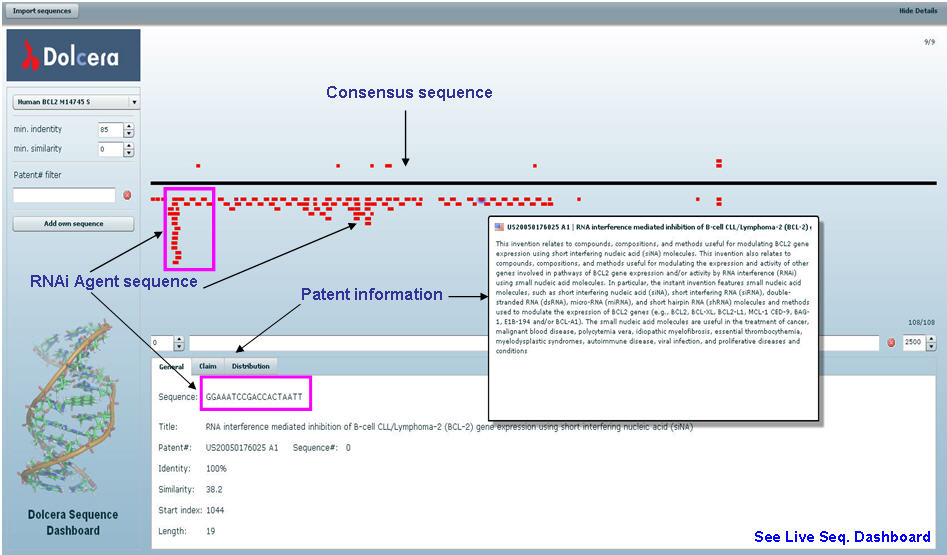
RNAi Sequence Dashboard
Sequence dashboard allows comparison of siRNA agents extracted from patents and scientific literature to the consensus sequence. Since not all the documents disclose the target sequence, the consensus sequence has to be identified and used for sequence dashboard. For example for Bcl-2, consensus sequence is retrieved from NCBI having seq. id M14745.
The dashboard also depicts the associated information with RNAi agent on the click of your mouse such as patent number, patent title, length of RNAi agent (siRNA), identity score etc.
Experimental data comparison matrix
Experimental data may be qualitative or quantitative, each being appropriate for different investigations.
Quantitative data was captured from following RNAi experiments for patent focusing on therapeutic application and compared.
- % inhibition of target mRNA expression
- % encapsulation efficiency
- % transfection efficiency
- % tumor volume reduction
- % cell count reduction
More such comparison matrix can be prepared for patent having varied focus such as on delivery vehicle, nature of RNAi agents, target gene source, system used to test RNAi etc.
Patent Focus- Therapeutic Application
| Sr. No. | Patent/Publication No. | Target gene | Experimental Details | RANK | |||||||
| BCL-2 | Others | Test Model | Inhibition of Target mRNA Expression (%) | Encapsulation Efficiency | Transfection efficiency Increase (%) | Tumor Volume Reduction (%) | Cell Count Reduction (%) | Other Remarks | |||
| 1 | US20070298056A1 | Yes | Human colon cancer (Caco-2) xenografts in nude mice | No | No | No | 90.08% Day 25 (Fig 2) |
No | No | 1 | |
| 2 | US20080038296A1 | Yes | In vivo (Mice) | No | No | No | 61.29% Day 13 (Fig 3) |
No | No | 2 | |
| 3 | US20070258952A1 | Yes | Human pancreatic carcinoma H79 xenografts | No | No | No | 49.23% Day 25 (Fig 4) |
No | No | 2 | |
| 4 | US20070172847A1 | Yes | Ramos cells | 80% (Fig 4. B) |
No | No | No | No | No | 2 | |
Ranking of patents can be done on the basis of focus of the study, experimental data disclosed, effectiveness of RNAi agent, type of model used to test the RNAi agent, etc.
Advantages:
- Information disclosure wise sorting available.
- Efficiency of different RNAi agent against target gene can be compared.
- Different models used to test cell line and the efficiency can also be compared.
Qualitative Data from Patent Documents
Experiments yielding qualitative data, mentioned below, can’t be compared but are equally important. We did not miss them!
- Gel electrophoresis
- Gel blots
- Tissue section* Cell staining
- RNAi agent construction
| Patent/ Publication No. | Gel blots | Gel Electrophorosis | Tissue section micrograph | Cell staining micrograph | RNAi Agent Construction |
| US20080021205A1 | Yes | _ _ | _ _ | Yes | _ _ |
| US20070275465A1 | _ _ | Yes | _ _ | _ _ | _ _ |
| WO2007013575A2 | _ _ | Yes | Yes | _ _ | _ _ |
| US20070081982A1 | _ _ | _ _ | _ _ | _ _ | Yes |
| WO2007013575A2 | Yes | _ _ | Yes | _ _ | _ _ |
| US20050266561A1 | _ _ | _ _ | _ _ | _ _ | Yes |